Amidst the ongoing opioid crisis, the number of individuals with opioid use disorder is on the rise. Opioid use disorder, a chronic, life-limiting illness, leads to substantial impairment and distress. As the population with opioid use disorder ages, there is a growing need for access to palliative care at the end of their lives.
This study led by Dr. Jenny Lau, the medical director of the Harold and Shirley Lederman Palliative Care Centre at Princess Margaret Cancer Centre, delves into the characteristics of people with opioid use disorder at the end of their lives and investigates the accessibility of palliative care for this demographic.
The research, based on a comprehensive study including information from ICES on 679,840 deaths, revealed that 11,200 of the deceased (1.6%) had opioid use disorder. Individuals with opioid use disorder died at younger ages than those without the disorder (50 years v. 78 years) and were more likely to reside in marginalized neighborhoods.
Comparatively, individuals with opioid use disorder were 16% less likely to receive palliative care than those without the disorder, particular in clinics and at home. This disparity is most likely linked to the high number of individuals with opioid use disorder who die suddenly from drug poisoning and do not have an opportunity to receive palliative care. The most common reasons why physicians provided palliative care for individuals with opioid use disorder were cancer, liver cirrhosis, and sepsis.
Dr. Lau, the lead author of this study, highlights, “The majority of conversations about the opioid crisis focus on the high number of opioid toxicity deaths. The unfortunate reality is that people with opioid use disorder are dying young from other causes as well. If we can improve our understanding of this population and the healthcare that they receive at the end of their lives, we can identify opportunities to intervene and improve their quality of life.”
This research underscores the urgency of addressing the disparities in palliative care provision for individuals with opioid use disorder. By understanding these discrepancies, researchers can pave the way for targeted interventions to ensure equitable end-of-life care for all.
With the proper training in both palliative care and addiction medicine and attention to socioeconomic inequalities, healthcare providers can significantly improve the quality of life for patients with opioid use disorder in their final days.

(L-R) Co-principal Investigators: Dr. Jenny Lau, the medical director of the Harold and Shirley Lederman Palliative Care Centre at Princess Margaret Cancer Centre, and Dr. Sarina Isenberg, chair in mixed methods palliative care research at Bruyère Research Institute, and adjunct scientist, ICES, Ottawa, Ontario.
This work was supported by funding from ICES, which is funded by a grant from the Government of Ontario. This work was also supported by the Ontario Health Data Platform (OHDP), a Province of Ontario initiative to support Ontario’s ongoing response to COVID-19 and its related impacts, and The Princess Margaret Cancer Foundation. This study also received funding from the Canadian Institutes of Health Research and Health Canada’s Health Care Policy and Strategies Program. The views expressed herein do not necessarily represent the views of Health Canada and Canadian Institutes of Health Research.
Jenny Lau reports honoraria from the University of Toronto and travel funding from the Canadian Institutes of Health Research and Health Canada. She is a member of the Global Institute of Psychosocial, Palliative and End-of-Life Care Operations Committee and medical director of the Harold and Shirley Lederman Palliative Care Centre. Additional information on competing interests can be found in the CMAJ publication.
Tara Gomes reports research funding from the Ontario Ministry of Health, the Ontario College of Pharmacists, and the Canadian Agency for Drugs and Technologies in Health; consulting fees from the Auditor General of British Columbia; payment for expert testimony from the Office of the Chief Coroner of Ontario; and travel support from Indigenous Services Canada. She is a Tier 2 Canada Research Chair in Drug Policy Research & Evaluation.
Sarina Isenberg reports travel funding from the American Association of Hospice Palliative Medicine. She is director of the St. Joseph’s Villa Foundation Board of Directors and a member of the Temple Anshe Sholom Succession Planning Committee
Lau J, Scott M, Everett K, Gomes T, Tanuseputro P, Jennings S, Bagnarol R, Zimmermann C, Isenberg S. Association between opioid use disorder and palliative care: a cohort study using linked health administrative data in Ontario, Canada. CMAJ. 2024 April 29. doi: 10.1503/cmaj.231419.
A research team at Toronto General Hospital Research Institute (TGHRI) has built an artificial intelligence (AI) learning model to understand complex biological interactions from large-scale datasets of the analysis of single cells.
Recent advancements in the study of the genes and gene expression patterns in single cells have provided a wealth of data that enables researchers to learn about cellular diversity, function, and how cells respond to various conditions. The use of a technique called single-cell RNA sequencing—a method that measures the levels of gene expression in each cell to determine how it functions—has led to the development of comprehensive data atlases.
“The large volume of sequencing data has created huge analytical challenges,” says Dr. Bo Wang, Scientist at TGHRI and senior author of the study. “To address this, we wanted to develop a foundation model to employ machine learning to decode and predict single-cell behaviours from sequencing data.”
A foundation model can be described as a giant database of information that is trained on a large number of diverse datasets and can be adapted for a variety of tasks. Language models, like chatGPT, are trained on text to learn patterns and meanings in language. Then, the model can be used to assist with tasks such as answering questions, summarizing text, or translating languages.
“While texts are made up of words, cells can be characterized by genes and the protein products they encode,” says Haotian Cui, doctoral student in Dr. Wang’s lab and co-first author of the study. “Using this principle, we developed a foundation model called scGPT (singe cell GPT) to examine single cell biology by pre-training on over 33 million cells.”
By training on a diverse dataset containing millions of cells from different tissues and conditions (i.e., cell types from 51 organs or tissues and 441 studies), scGPT has learned to understand patterns in gene expression and cell behavior and has been taught to create new information based on what it learned. Its main part uses special tools called transformer blocks to help it understand and process the data. After its initial training, its settings can be adjusted to make it work better with new information, which can be useful for various tasks.
The team found that scGPT is effective for tasks such as identifying cell types, predicting gene activity in cells, correcting batch effect errors in sequencing data, and uncovering important gene interactions that vary depending on the cell type or condition. This approach enhances the modeling of single-cell sequencing data and provides valuable insights into gene-gene interactions specific to different conditions such as cell states and gene expression disruptions.
“The release of scGPT models and workflows will be able to accelerate research in cellular biology and beyond, offering a standardized approach for analyzing single-cell omics—the profiling of single cells in various populations,” says Chloe Wang, co-first author of the study and doctoral student at TGHRI.
By leveraging the power of a pre-trained generative AI model, researchers hope to pave the way for innovative therapeutic strategies and deepen understanding of cellular processes.
“For the future, our goal is to make our model smarter and better at understanding how cells work in different situations,” adds Dr. Wang who is also Chief AI Scientist at UHN and co-lead of the UHN AI Hub.
Since the preprint of this study in May 2023 and the release of scGPT, it has significantly impacted the field, with over 13,000 installations and 55 citations before its official publication.
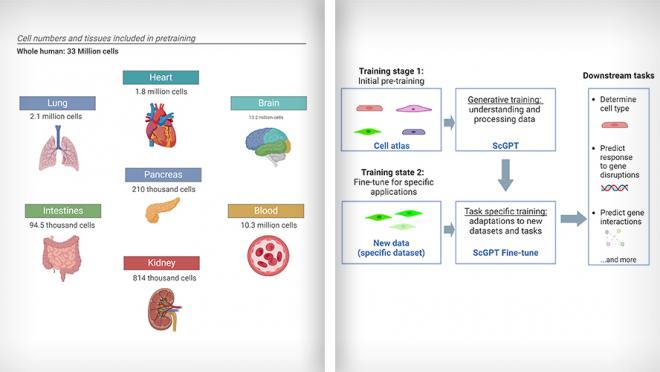
(L) scGPT was trained using single cell RNA-sequencing data from 33 million normal human cells from various organs. (R) Simplified workflow of scGPT. Images adapted from Cui et al., 2024, Nature Methods using BioRender.
This work was supported by funding from the Natural Sciences and Engineering Research Council of Canada, the Canadian Institute For Advanced Research AI Chairs Program, and the Peter Munk Cardiac Centre at the University Health Network.
Dr. Bo Wang is a Tier 2 Canada Research Chair in Artificial Intelligence for Medicine and an Assistant Professor at the University of Toronto.
Dr. Bo Wang is on the advisory board of Vevo Therapeutics and co-author Dr. Nan Duan is an employee of Microsoft and holds equity in the company.
Cui H, Wang C, Maan H, Pang K, Luo F, Duan N, Wang B. scGPT: toward building a foundation model for single-cell multi-omics using generative AI. Nat Methods. 2024 Feb 26. doi: 10.1038/s41592-024-02201-0. Epub ahead of print. PMID: 38409223.
Krembil’s annual Research Day is a unique opportunity to exchange ideas, foster collaboration and celebrate research achievements.
This year’s event, held at the MaRS Centre on Thursday, April 11, 2024, welcomed 230 trainee researchers, faculty and staff who showcased the breadth of talent within our Institute and our collective dedication to advancing scientific knowledge.
The event opened with a welcome video that shined a spotlight on what it is like to be a Krembil trainee, as well as opening remarks from Dr. Jaideep Bains, Director of the Krembil Research Institute. Dr. Bains commended trainees at all levels for their diverse talents, dedication to research and outstanding contributions to the Institute.
Throughout the event, nine trainees delivered oral presentations that highlighted achievements across the Institute’s three research pillars: brain and spine, vision, and bone and joints. The event also featured more than 70 poster presentations from trainees and research staff that facilitated scientific dialogue and idea sharing.
This year's Research Day also featured a special “Lunch and Learn” event. Internationally renowned communications expert Bri McWhorter, Founder and CEO of Activate to Captivate, shared valuable insights on science communication in a talk titled, “The Art of Delivering an Unforgettable Presentation.”
Trainees benefited from McWhorter's expertise in storytelling, learning strategies for creating connections with audiences, navigating key moments in talks and crafting engaging narratives.
The event concluded with a keynote address from Dr. Anne-Marie Malfait, Professor of Internal Medicine and The George W. Stuppy, MD, Chair of Arthritis at Rush University, titled “Chronic Joint Pain: Five Short Lessons from Osteoarthritis.”
Click here to watch a short slideshow recapping this year's Research Day.
“Thank you to everyone who joined us for our annual Research Day. Your participation made the event an inspiring celebration of the breadth of ideas and expertise at our Institute and the collaborative spirit that fuels our success,” said Dr. Bains. “I look forward to carrying this momentum forward as we plan for next year’s Research Day and more opportunities for our research community to come together.”
The Krembil community thanks the many individuals who made this year’s Research Day possible, including the Krembil Trainee Affairs Committee—chaired by Dr. Mary Pat McAndrews—the Krembil Communications and Administration teams, the oral presentation session Chairs—Drs. Joan Wither and Bill Hutchison—and everyone who served as judges for the oral and poster presentations.
Presentation Awards
The following trainees and postdoctoral researchers received awards for best oral and poster presentations:
Poster Presentations - Graduate Student Category:
● 1st place: Zi Xuan Zhang
● 2nd place (tied): Ain Kim
● 2nd place (tied): Anca Maglaviceanu
Poster Presentations - Postdoctoral Researcher Category:
● 1st place: Laura Whitall-Garcia
● 2nd place: Icaro Oliveira
● 3rd place: Cody Wilson-Konderka
Oral Presentations - Graduate Student Category:
● 1st place: Kabriya Thavaratnam
● 2nd place (tied): Brian Nghiem
● 2nd place (tied): Pedram Mouseli
Oral Presentations - Postdoctoral Researcher Category:
● 1st place: Emily Mills
A notable addition to this year's awards was the People's Choice Award for Oral Presentations, received by doctoral student Kabriya Thavaratnam for her exceptional research and presentation skills. Judges also awarded master’s student Nikou Kelardashti for her sex and gender-informed abstract, reflecting Krembil's commitment to promoting diversity and inclusion in research.
Congratulations to the winners and to everyone who presented their work. We look forward to seeing you at next year’s Research Day!
Researchers at UHN's McEwen Stem Cell Institute shed light on the role of cardiac fibroblasts in heart development and disease, key cells for improving treatments for heart-related conditions, specifically those affecting the heart muscle.
Cardiac fibroblasts make up approximately 15–30% of all heart cells and are essential for both fetal heart development and maintenance of the adult heart structure. The majority of the fibroblast population is derived from the outer layer of the heart, called the epicardium.
"There's a lack of laboratory models that accurately replicate the formation and function of the epicardium, which hinders our understanding of heart development, disease mechanisms, and the advancement of new therapies," explains Dr. Gordon Keller, Director of the McEwen Stem Cell Institute and senior author of this study.
To address this gap, the team developed cardiac organoids, three-dimensional pluripotent stem cell-derived structures, designed to mimic key aspects of human heart development and function. These organoids spontaneously form the functional muscle tissue and epicardium found in the heart, providing a platform to study cardiac cell interactions in a more physiologically relevant environment.
Using advanced techniques to monitor cell differentiation and maturation, researchers discovered that interactions between epicardial cells and heart muscle cells (cardiomyocytes) within the organoids resulted in the specification and migration of fibroblasts, in a pattern similar to that observed in developing human hearts.
"Our study demonstrates that this organoid platform serves as an advanced model system for studying multicellular mechanisms underlying heart disease," explains Dr. Ian Fernandes Scientific Associate at McEwen Stem Cell Institute and first author of this study.
Moreover, single-cell RNA sequencing, a cutting-edge technique that allows scientists to study the molecular profile of individual cells, revealed significant diversity within the populations of fibroblasts and cardiomyocytes in the cardiac organoids. This diversity resembled what is seen in both healthy and diseased adult hearts.
Researchers also identified distinct subpopulations of fibroblasts, including a group expressing the CD9 protein, which has been associated with reparative functions.
"The identification of distinct subpopulations of cells within the heart organoids, especially those with reparative properties like the CD9+ fibroblasts, opens up new avenues for understanding and potentially treating heart diseases," adds Dr. Keller. "By targeting these specific cell types, we may develop more precise and effective therapies for various heart-related conditions."
This research opens avenues for understanding cardiac development, disease mechanisms, and potential regenerative therapies, promising advancements in heart-related treatments. Additionally, the ability to isolate and co-transplant specific fibroblast subpopulations with cardiomyocytes in damaged hearts could hold potential for generating new heart muscle and improving heart function.
This work was supported by the Canadian Institutes of Health Research and UHN Foundation. Dr. Gordon Keller is a Professor at the Department of Medical Biophysics at the University of Toronto.
G.M.K. is a scientific co-founder and paid consultant for BlueRock Therapeutics LP and a paid consultant for VistaGen Therapeutics.
Fernandes, I., Funakoshi, S., Hamidzada, H. et al. Modeling cardiac fibroblast heterogeneity from human pluripotent stem cell-derived epicardial cells. Nat Commun 14, 8183 (2023). https://doi.org/10.1038/s41467-023-43312-0
Researchers from UHN’s KITE Research Institute offer new hope for epilepsy research as they develop deep-learning models to predict epileptic seizures.
Epilepsy, one of the world’s most prevalent neurological disorders, affects over 50 million people worldwide. Characterized by the sudden onset of seizures, epilepsy can lead to serious physical injury and even death. The ability to predict the onset of epileptic seizures can significantly reduce injury and improve quality of life.
A team led by Dr. Shehroz Khan, KITE Scientist and senior author of the study, focused on leveraging deep learning models to analyze electroencephalogram (EEG) data. EEG is a test that uses small electrodes to measure brain activity and serves as a vital tool for understanding seizure onset.
“Deep learning models are advanced computer algorithms that learn to recognize patterns and make predictions by processing large amounts of complex data. By using these models to distinguish pre-seizure EEG patterns, we can help epilepsy patients and their caregivers anticipate seizures and take preventive measures,” states Dr. Khan.
Using a combination of supervised and unsupervised deep learning approaches, the researchers trained the learning models to identify subtle changes in brain activity preceding seizures.
“Supervised deep learning involves using labelled data where seizure occurrence is known. On the other hand, unsupervised deep learning allows the model to learn predictive patterns from unlabeled data on its own,” explains Zakary Georgis-Yap, a previous master's student in Dr. Khan’s lab and first author of the study. “The advantage of unsupervised learning models is that they do not require comprehensively labelled data—which can be challenging and time-consuming to obtain.”
To evaluate the effectiveness of their models, the researchers conducted extensive testing on two large seizure datasets containing EEG-recorded data from 40 patients.
The results of the study were promising, showcasing the feasibility of both supervised and unsupervised approaches in seizure prediction. However, prediction results for both models varied across datasets, patients, and learning approaches, highlighting the considerable variability in pre-seizure brain activity between individuals.
“While there is still work to be done, our research represents a significant step forward in the field of epilepsy management,” concludes Dr. Khan. “By harnessing the potential of deep learning, we have the opportunity to develop personalized therapeutic interventions and ultimately save lives.”
This work was supported by the Natural Sciences and Engineering Research Council of Canada, the Data Science Institute at the University of Toronto and UHN Foundation. Dr. Shehroz Khan is an Assistant Professor at the University of Toronto’s Institute of Biomedical Engineering.
#Georgis-Yap, Z., Popovic, M.R. & #Khan, S.S. Supervised and Unsupervised Deep Learning Approaches for EEG Seizure Prediction. J Healthc Inform Res. 2024 Jan 4. DOI: 10.1007/s41666-024-00160-x
#Zakary Georgis-Yap and Dr. Shehroz Khan contributed equally to the study.
Researchers from Princess Margaret Cancer Centre (PM) have developed an approach to predict outcomes in patients treated with pembrolizumab—an immunotherapy drug used to treat various cancers such as head and neck, breast, and skin cancer.
Analysis for this study was performed using blood samples collected from patients with different cancer types treated with pembrolizumab in the INSPIRE trial, an investigator-initiated study that was designed and carried out at PM.
Pembrolizumab is a type of antibody that attaches to a protein known as programmed death 1 (PD-1), hindering its activity. PD-1 acts as a checkpoint for the immune system, preventing it from becoming overactive and attacking healthy cells. But in the case of cancer, PD-1 can also act as a brake to prevent the immune system from attacking cancer cells. Therefore, inhibiting PD-1 releases the brakes on the immune system and allows it to recognize and attack cancer cells more effectively.
“Although pembrolizumab is used to target many cancers, some patients may develop resistance to the drug,” says Dr. Stutheit-Zhao, Postdoctoral Researcher at PM and co-first author of the study. “Biomarkers like circulating tumour DNA (ctDNA) can provide insights into the effectiveness of treatments without the need for invasive procedures. However, many current ctDNA analyses are so-called tumour-informed, thus requiring molecular sequencing of patients' tumour tissue to identify alterations that can be tracked in the blood.”
ctDNA are pieces of tumour DNA that circulate in the bloodstream. Previous studies—including a clinical trial from this team—have shown that changes in levels of ctDNA using a tumour-informed assay early on during the treatment course can predict pembrolizumab treatment success and patient survival. However, as this assay also requires tumour samples, it may not be feasible for some patients as tumours may be in challenging locations to biopsy, or biopsy specimens may contain insufficient tumour cells for genetic analysis.
“We wanted to investigate a way to predict treatment outcome using ctDNA alone without needing to analyze a tumour biopsy,” says Dr. Enrique Sanz-Garcia, Clinician Investigator at PM and co-first author of the study. “DNA from tumours is known to be enriched for a type of modification, called methylation. Therefore, we analyzed methylated cell-free DNA from 204 plasma samples from 87 patients before and during treatment with pembrolizumab from the INSPIRE trial. This particular assay does not require access to patients’ tumour samples.”
Previous studies have demonstrated the ability of a technique, first introduced by scientists at UHN, that isolates and analyzes this methylated DNA, called cell-free methylated DNA immunoprecipitation and sequencing (cfMeDIP-seq), to estimate ctDNA levels. This technique can also quantify DNA fragment lengths, which are known to be shorter in tumour tissue compared to DNA derived from normal tissue.
The team therefore retrospectively tested whether methylation and fragmentation status of cell-free DNA using cfMeDIP-seq analysis can monitor the response to pembrolizumab in the INSPIRE study.
“Our findings indicated a strong correlation between the ctDNA analysis of methylation patterns and DNA fragmentation in blood samples alone with tumour-informed ctDNA analysis,” says Dr. Lilllian Siu, Senior Scientist at PM and senior author of the study. “Importantly, methylation analysis predicted overall survival and progression-free survival, and fragment length analysis was able to predict overall survival in patients treated with pembrolizumab.”
Using statistical analysis, the team found that a decrease in cancer-specific methylation over the first six weeks of pembrolizumab treatment was associated with a 60% lower risk of death and a 65% lower risk of disease progression compared to the reference group. For fragmentation analysis, a decrease in fragment length score over time was significantly associated with a 60% lower risk of death.
This suggests that analysis of circulating DNA assayed by cfMeDIP-seq yields promising tumour-naïve (i.e. not requiring access to tumour samples) predictive biomarkers for response to pembrolizumab.
“This is the first reported examination of methylated cell-free DNA in advanced cancer patients during treatment with pembrolizumab and one of the first direct comparisons between ctDNA analysis with and without tumour samples,” adds Dr. Trevor Pugh, Senior Scientist at PM and co-senior author of the study.
This study, published in Cancer Discovery, the lead journal of the American Association for Cancer Research, reveals the potential of a minimally invasive blood test (i.e., a liquid biopsy) for predicting response to cancer treatments. This could enable earlier response assessment by doctors and lead to prompt redirection to next-line treatment options in non-responders before radiological assessments.
Circulating tumour DNA (ctDNA) can be analyzed using blood tests as part of a liquid biopsy. Liquid biopsies are blood tests used to detect cancer cells or other markers of disease.
This work was supported by The Princess Margaret Cancer Foundation, the Ontario Institute for Cancer Research, the Terry Fox Research Institute, the Princess Margaret Cancer Centre Global Oncology Program, and the Cancer Research Institute.
Dr. Enrique Sanz-Garcia is an Assistant Professor at the University of Toronto (U of T). Dr. Lilian Siu is a Professor of Medicine at U of T and holds the BMO Chair in Precision Cancer Genomics. Dr. Pugh is a Professor in the Department of Medical Biophysics at U of T and is a tier 2 Canada Research Chair in Biological Physics.
Dr. Sanz Garcia reported research funding from Novartis. Dr. Siu reported either a leadership role, financial interest in, or a consulting role with Treadwell Therapeutics, Agios, Merck, AstraZeneca/MedImmune, Roche, Voronoi Health Analytics, Oncorus, GlaxoSmithKline, Seattle Genetics, Arvinas, Navire, Janpix, Relay Therapeutics, Daiichi Sankyo/UCB Japan, Janssen, Hookipa Pharma, InterRNA, Tessa Therapeutics, Sanofi, Amgen, and research funding from Bristol Myers Squibb (Inst),Genentech/Roche (Inst), GlaxoSmithKline (Inst), Merck (Inst), Novartis (Inst), Pfizer (Inst), AstraZeneca (Inst), Boehringer Ingelheim (Inst), Bayer (Inst), Amgen (Inst), Astellas Pharma (Inst), Shattuck Labs (Inst), Symphogen (Inst), AVID Radiopharmaceuticals (Inst), Mirati Therapeutics (Inst), Intensity Therapeutics (Inst), Karyopharm Therapeutics (Inst). Dr. Pugh has provided consultation for AstraZeneca, Chrysalis Biomedical Advisors, Merck, and SAGA Diagnostics (compensated); and receives research support (institutional) from Roche/Genentech.
For a full list of competing interests, see manuscript.
Stutheit-Zhao EY*, Sanz-Garcia E*, Liu ZA, Wong D, Marsh K, Abdul Razak AR, Spreafico A, Bedard PL, Hansen AR, Lheureux S, Torti D, Lam B, Yang SYC, Burgener J, Luo P, Zeng Y, Cheng N, Awadalla P, Bratman SV, Ohashi PS, Pugh TJ, Siu LL. Early changes in tumor-naive cell-free methylomes and fragmentomes predict outcomes in pembrolizumab-treated solid tumors. Cancer Discov. 2024 Feb 23. doi: 10.1158/2159-8290.CD-23-1060. Epub ahead of print. PMID: 38393391.
*authors contributed equally to this article.
Researchers from UHN’s KITE Research Institute have investigated a new affordable and clinically accessible training system for improving the standing balance of spinal cord injury patients.
Spinal cord injury affects approximately 85,000 Canadians every year. Individuals with incomplete spinal cord injury can regain their ability to walk; however, a majority of these individuals experience falls at least once a year. Falling can reduce mobility, physical activity, and also quality of life.
Balance therapy helps increase muscle strength and reaction but relies on an individual’s vision to help maintain standing balance. Visual feedback balance training (VFBT) is a training method that incorporates visual cues to help improve balance and postural control. It involves shifting your body toward a target location on a screen.
Functional electrical stimulation is an additional rehabilitation technique that uses electrical impulses to activate muscles, helping individuals regain movement and improve function. Researchers have explored combining VFBT with functional electrical stimulation as a comprehensive rehabilitation approach for improving balance control. However, these rehabilitation systems traditionally rely on costly force plates to measure a participant’s movement.
The team led by Dr. Kei Masani, KITE Senior Scientist and senior author of the study, investigated the integration of low-cost and portable sensors like a depth camera and pressure mat, which use motion tracking and distribution of pressure, respectively, to analyze movement.
The effectiveness of these sensors was put to the test by measuring the movements of ten able-bodied participants, with no history of neurological disorders, as they completed balance rehabilitation exercises using the combined VFBT and functional electrical stimulation system. Researchers found that the depth camera outperformed the pressure mat, showing higher accuracy and lower error relative to the force plate, in capturing crucial balance and movement measures.
These results suggest that the depth camera could replace the force plate in the VFBT and functional electrical stimulation rehabilitation system.
Derrick Lim, PhD candidate in the lab of Dr. Masani and first author of the study, is enthusiastic about these findings stating, “This study marks a significant step to making rehabilitation more accessible and effective for individuals with spinal cord injuries. The use of affordable sensors opens doors to broader implementation in clinical settings, ultimately improving the quality of life for patients.”
Future work will focus on using this system with individuals who have experienced spinal cord injury and have poorer balance capabilities. It has potential applications in other populations who experience neurological damage resulting in balance impairments, such as adults living with stroke.
This work was supported through the Collaborative Health Research Projects program, a joint initiative between both, the Canadian Institutes of Health Research (CIHR) and the Natural Sciences and Engineering Research Council of Canada (NSERC), as well as UHN Foundation. Dr. Kei Masani is an Associate Professor at the Institute of Biomedical Engineering at the University of Toronto.
Lim D, Pei W, Lee JW, Musselman KE, Masani K. Feasibility of using a depth camera or pressure mat for visual feedback balance training with functional electrical stimulation. Biomed Eng Online. 2024 Feb 12. doi: 10.1186/s12938-023-01191-y.
Research conducted at UHN's research institutes spans the full spectrum of diseases and disciplines, including cancer, cardiovascular sciences, transplantation, neural and sensory sciences, musculoskeletal health, rehabilitation sciences, and community and population health.
Learn more about our institutes by clicking below: